Calculate richness estimate and add to phyloseq sample data
Source:R/ps_calc_richness.R
ps_calc_richness.RdWrapper around microbiome::richness() function. Takes and returns a phyloseq object. Calculates a richness estimate at a given taxonomic rank. Returns phyloseq unaggregated, with an additional variable. Variable name is by default created by pasting the index and rank.
Usage
ps_calc_richness(
ps,
rank,
index = "observed",
detection = 0,
varname = paste0(index, "_", rank)
)Details
Don't filter taxa before calculating richness.
These richness indices are estimates. For a discussion of the uncertainty and bias of these estimates see e.g. work by Amy Willis https://doi.org/10.3389/fmicb.2019.02407
See also
microbiome::richness
Examples
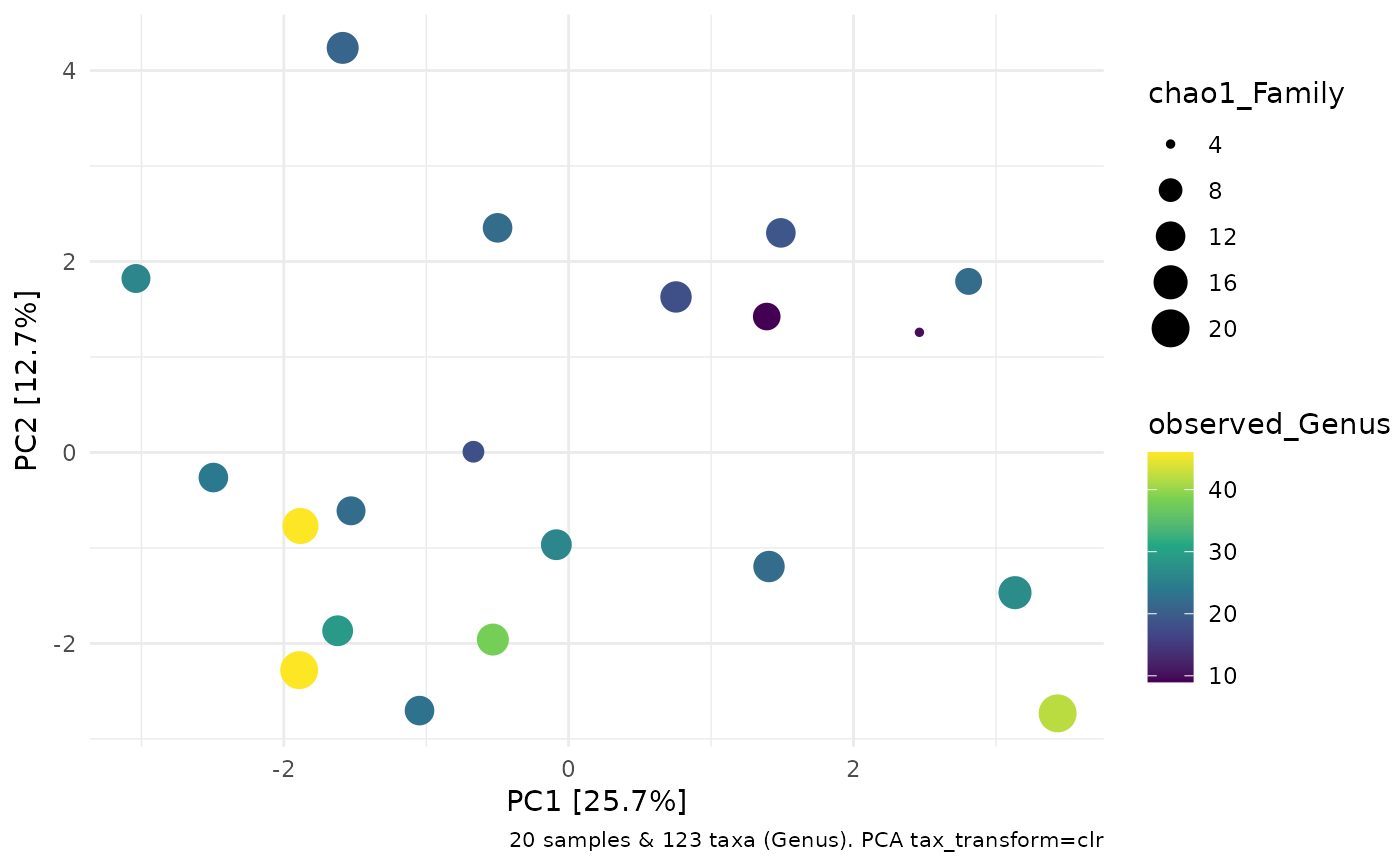
data(ibd, package = "microViz")
ibd %>%
ps_filter(abx == "abx") %>%
tax_fix() %>%
ps_calc_richness("Genus", index = "observed") %>%
ps_calc_richness("Family", index = "chao1") %>%
tax_transform(rank = "Genus", transform = "clr") %>%
ord_calc("PCA") %>%
ord_plot(
colour = "observed_Genus", size = "chao1_Family"
) +
ggplot2::scale_colour_viridis_c()
#> Warning: `transformation` argument deprecated, use `trans` instead.